Research

Welcome to the Children's Neuroscience Institute (CNI), the destination for pediatric neuroscience research, education, and patient care.
One of the most common causes of death and disability in US children is brain injury. It can be extremely challenging to accurately diagnose and provide prognostication of neurological diseases in children. Currently, translation of therapies from laboratory to patients takes a very long time, thus, transformative approaches are desperately needed for the development of individualized treatments and accelerate recovery.
Our purpose is to improve child health and combat childhood neurological diseases by catalyzing multidisciplinary research collaborations across the neuroscience community. We want to build a unique and highly successful program that will generate innovative neuroscience methods, diagnostics, and treatments; making the CNI the top destination for treatment, education, and science.
Udai Pandey, PhD
Associate Professor
Pediatrics (Primary) and Human Genetics (Secondary)
Director, Children's Neuroscience Institute
Multiomics
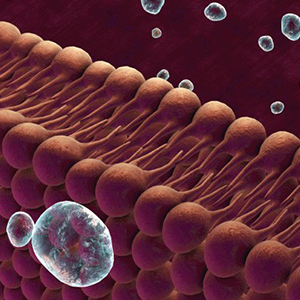 Lipids are important chemical building blocks in the human body, particularly in the central nervous system (CNS) where the lipid content is the highest next to adipose tissue. Apart from roles as chemical building blocks, and perhaps more importantly, lipids function as signaling molecules in the regulation of cellular life span, inflammation, and immunity. Various lipidomic approaches are utilized for the identification and deciphering of the signaling roles of lipids.
Lipids are important chemical building blocks in the human body, particularly in the central nervous system (CNS) where the lipid content is the highest next to adipose tissue. Apart from roles as chemical building blocks, and perhaps more importantly, lipids function as signaling molecules in the regulation of cellular life span, inflammation, and immunity. Various lipidomic approaches are utilized for the identification and deciphering of the signaling roles of lipids.
A number of lipidomic approaches have been pioneered at the University of Pittsburgh, among them is oxidative lipidomics that focuses on oxidized lipids and analysis of single cell lipidome. Using these newly developed methods, specific oxidized lipids have been identified as cellular death signals, mediators of tumor immunity and inflammation.
These methods are also identified the brain mitochondria (the powerhouse of cells) as having a specific complement of lipids that are different and distinct from all other organs. When the brain is injured, these mitochondria-specific lipids are released into the blood, thus having the ability to aid in diagnosis and prognostication. Imaging lipids at a single-cell level or even subcellular level has become possible only recently. Approaches have been developed using mass spectrometry to detect and explore the localization of lipids in cells. A number of lipidomic approaches have been pioneered at the University of Pittsburgh, among them is oxidative lipidomics that focuses on oxidized lipids and analysis of single-cell lipidome. Using these newly developed methods, specific oxidized lipids have been identified as cellular death signals, mediators of tumor immunity and inflammation.
We envision coupling of the single cell lipidomics maps with protein and DNA maps to obtain a complete understanding of cellular metabolism and signaling. We believe this knowledge will have a transformative impact in diagnostic and therapeutic avenues for CNS disorders.
iPSC
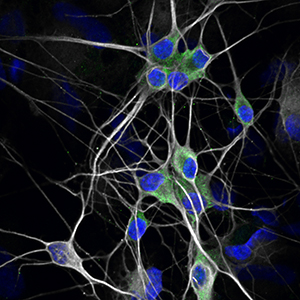 This theme will use the latest techniques to reprogram, expand, and characterize human induced pluripotent stem cells (iPSC) from both healthy and diseased skin or blood tissue. The iPSC will then be turned into specific human cells, including specific components of the nervous system and other tissue cells.
This theme will use the latest techniques to reprogram, expand, and characterize human induced pluripotent stem cells (iPSC) from both healthy and diseased skin or blood tissue. The iPSC will then be turned into specific human cells, including specific components of the nervous system and other tissue cells.
Some applications of this technology include human diseases ‘modeling-in-a-dish’, developing human reporter cell lines via genetic modification, drug screening on pathological human cell types, and potentially developing cell replacement or regenerative therapies.
Udai Bhan Pandey, PhD
Associate Professor, Pediatrics and Human Genetics and Neurology, University of Pittsburgh School of Medicine
Bioinformatics
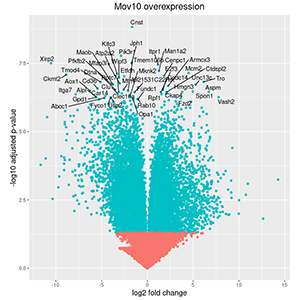 Next-generation sequencing, and other high-throughput technologies have revolutionized the field of medicine with the ability to personalize diagnosis, risk assessment and treatment of patients. Given the massive amount of data generated, there is a greater need for computational resources and tools to analyze, interpret, visualize, and store the information.
Next-generation sequencing, and other high-throughput technologies have revolutionized the field of medicine with the ability to personalize diagnosis, risk assessment and treatment of patients. Given the massive amount of data generated, there is a greater need for computational resources and tools to analyze, interpret, visualize, and store the information.
As an interdisciplinary science, bioinformatics leverages mathematics, statistics, and computer science to analyze and interpret biological information at the molecular, cellular, and genomic level. Bioinformatics has a profound impact in the better understanding of disease mechanisms, the identification of biomarkers of particular diseases or conditions, characterization of the patients' drug responses, and expediting the progress towards precision medicine.
The Bioinformatics Core is a resource to support the bioinformatics needs of investigators for sequencing datasets generated in house by the HSSC@CHP core at Rangos Research Center, as well as external sources. The essence of a bioinformatics core is to provide a variety of data analysis services, but she also believes in empowering others to be able to do their own analysis through training events and workshops.
Dhivyaa Rajasundaram, PhD
Research Assistant Professor, Department of Pediatrics
Lead, Bioinformatics Core, UPMC Children's Hospital of Pittsburgh
Molecular Imaging
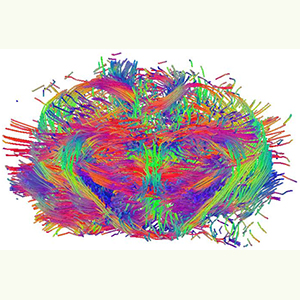 Using high-tech software and machinery, the imaging theme creates two- and three-dimensional images in addition to longitudinal quantification. Molecular images provide visualization, characterization and measurements of biological tissue and other living specimens at the cellular and molecular levels.
Using high-tech software and machinery, the imaging theme creates two- and three-dimensional images in addition to longitudinal quantification. Molecular images provide visualization, characterization and measurements of biological tissue and other living specimens at the cellular and molecular levels.
The Molecular Imaging Core provides the following services: (1) consultation for design of and implementation of in vivo organ/tissue imaging and image-guided interventions as part of investigator-initiated research grants or of inter-/intra-institutional program grants; (2) support to investigators in image acquisition and image-guided interventions; (3) assist in image process and data analysis; and (4) maintain instruments and provide standard supplies for the imaging.
Yijen Wu, PhD
Assistant Professor, Department of Developmental Biology
Director, Animal Imaging Core, UPMC Children's Hospital of Pittsburgh
Human Neurocognitive Development
 Cognition and brain systems develop significantly from the newborn period through adolescence and into adulthood, establishing lifelong trajectories. Understanding the brain mechanisms that support neurocognitive development can inform how to detect risk for impaired development (e.g., psychopathology, sequelae from traumatic brain injury, stress, or illness) and approaches for supporting optimal development (e.g., cognitive therapy, training, pharmaceuticals, brain stimulation). Neuroimaging approaches including: task and resting state functional Magnetic Resonance Imaging (t- rs- fMRI), brain structural MRI (sMRI), Magnetic Resonance Spectroscopy (MRS), Diffusion Tensor Imaging (DTI), Electroencephalography (EEG), Magnetoencephalography (MEG), Functional near-infrared spectroscopy (fNIRS), and Positron Emission Tomography (PET) are used to map changes in brain structure, function, and neurochemistry that support cognitive development. Identifying normative development will establish a pediatric neurocognitive growth chart needed to assess the impact of impairments or trauma, build predictive models of recovery, and assess recovery progress enhancing life trajectories.
Cognition and brain systems develop significantly from the newborn period through adolescence and into adulthood, establishing lifelong trajectories. Understanding the brain mechanisms that support neurocognitive development can inform how to detect risk for impaired development (e.g., psychopathology, sequelae from traumatic brain injury, stress, or illness) and approaches for supporting optimal development (e.g., cognitive therapy, training, pharmaceuticals, brain stimulation). Neuroimaging approaches including: task and resting state functional Magnetic Resonance Imaging (t- rs- fMRI), brain structural MRI (sMRI), Magnetic Resonance Spectroscopy (MRS), Diffusion Tensor Imaging (DTI), Electroencephalography (EEG), Magnetoencephalography (MEG), Functional near-infrared spectroscopy (fNIRS), and Positron Emission Tomography (PET) are used to map changes in brain structure, function, and neurochemistry that support cognitive development. Identifying normative development will establish a pediatric neurocognitive growth chart needed to assess the impact of impairments or trauma, build predictive models of recovery, and assess recovery progress enhancing life trajectories.
Beatriz Luna, PhD
Staunton Professor of Psychiatry and Pediatrics
Professor of Psychology at the University of Pittsburgh
Director, Laboratory of Neurocognitive Development
Editor in Chief, Journal Developmental Cognitive Neuroscience
Founder and President, Flux Society for Developmental Cognitive Neuroscience
